
Health & Medicine
Why are there so few drugs to treat viruses?
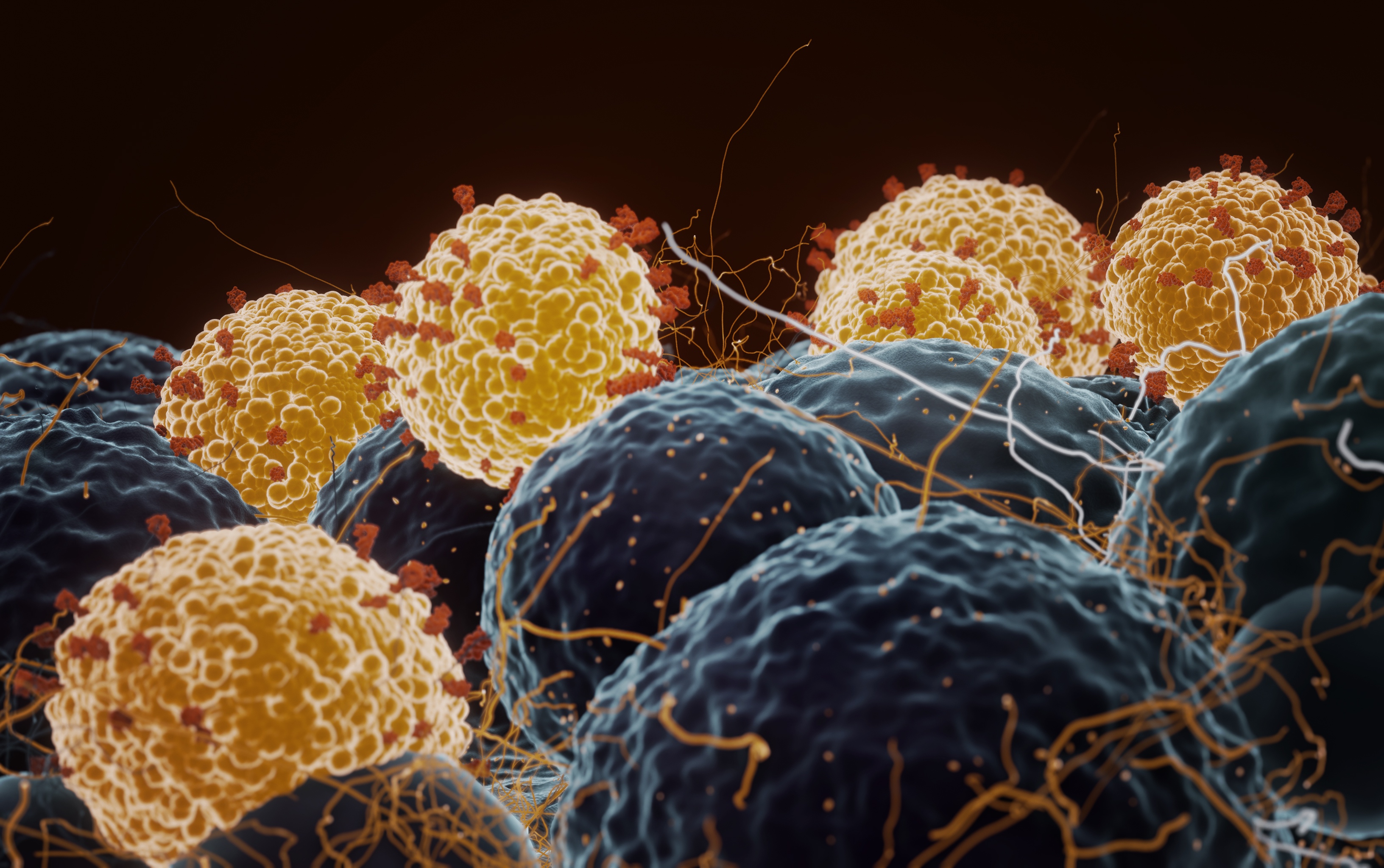
Many viruses – including coronaviruses – have a protective outer layer of proteins, fats and sugars. New research shows targeting sugar production has potential for broad-spectrum antiviral drugs
Published 9 November 2020
Despite its ability to wreak havoc across the globe, the SARS-CoV-2 coronavirus isn’t alive. It is merely a ribbon of genetic material, encased in an envelope of proteins, fats and sugars.
A virus’ primary goal is to replicate– make copies of itself – but on their own viruses can do nothing. They need a host, be it bat, bird or human.
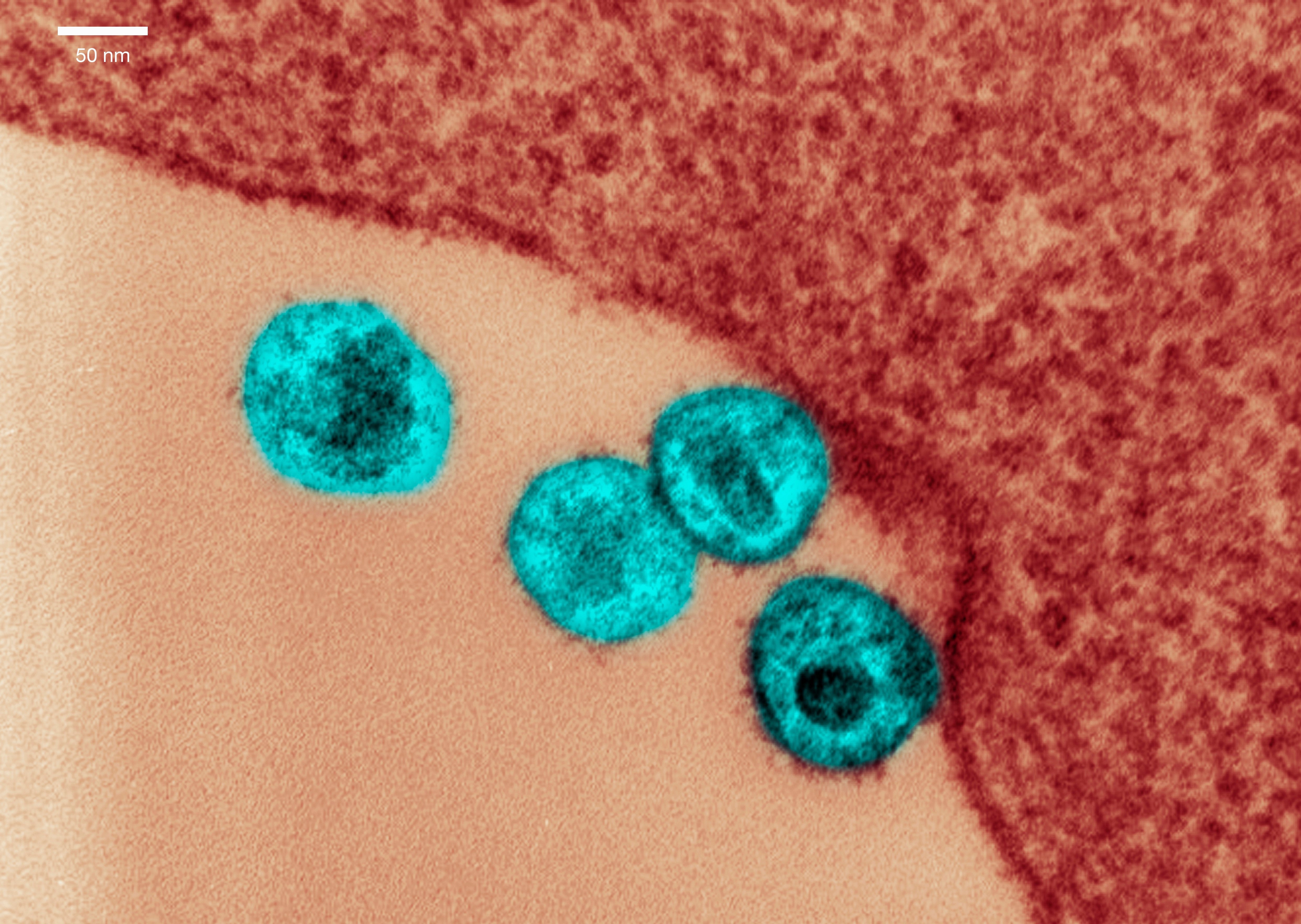
After infecting the host, the virus’ genetic information – which in the case of SARS-CoV-2 isn’t DNA but RNA – escapes from its surrounding layers, hijacking the host cell’s machinery to build the proteins it needs to replicate.

Health & Medicine
Why are there so few drugs to treat viruses?
So how do we stop an enemy that isn’t alive, but hides within our cells? This is a question asked by many scientists across the globe, with answers urgently needed for many viruses including SARS-CoV2.
Professor Spencer Williams from the University of Melbourne, and Professor Gideon Davies from the University of York in the UK, along with colleagues from the Universities of Warwick and Oxford, may have found a way to derail viral replication that could in turn become a broad spectrum anti-viral drug target.
“All viruses are made up of a genetic code inside a protein coat known as a capsid,” explains Professor Williams.
“Many viruses including hepatitis B, dengue, HIV, SARS-CoV-2 are also surrounded by a protective second layer called an envelope, giving them the label ‘encapsulated viruses’. This second layer is made of proteins, fats and sugars, and is the target of our antiviral drug work”
Every living thing contains some form of protein. They are versatile molecules that carry out a multitude of biochemical reactions like energy creation, and providing structure to cells.
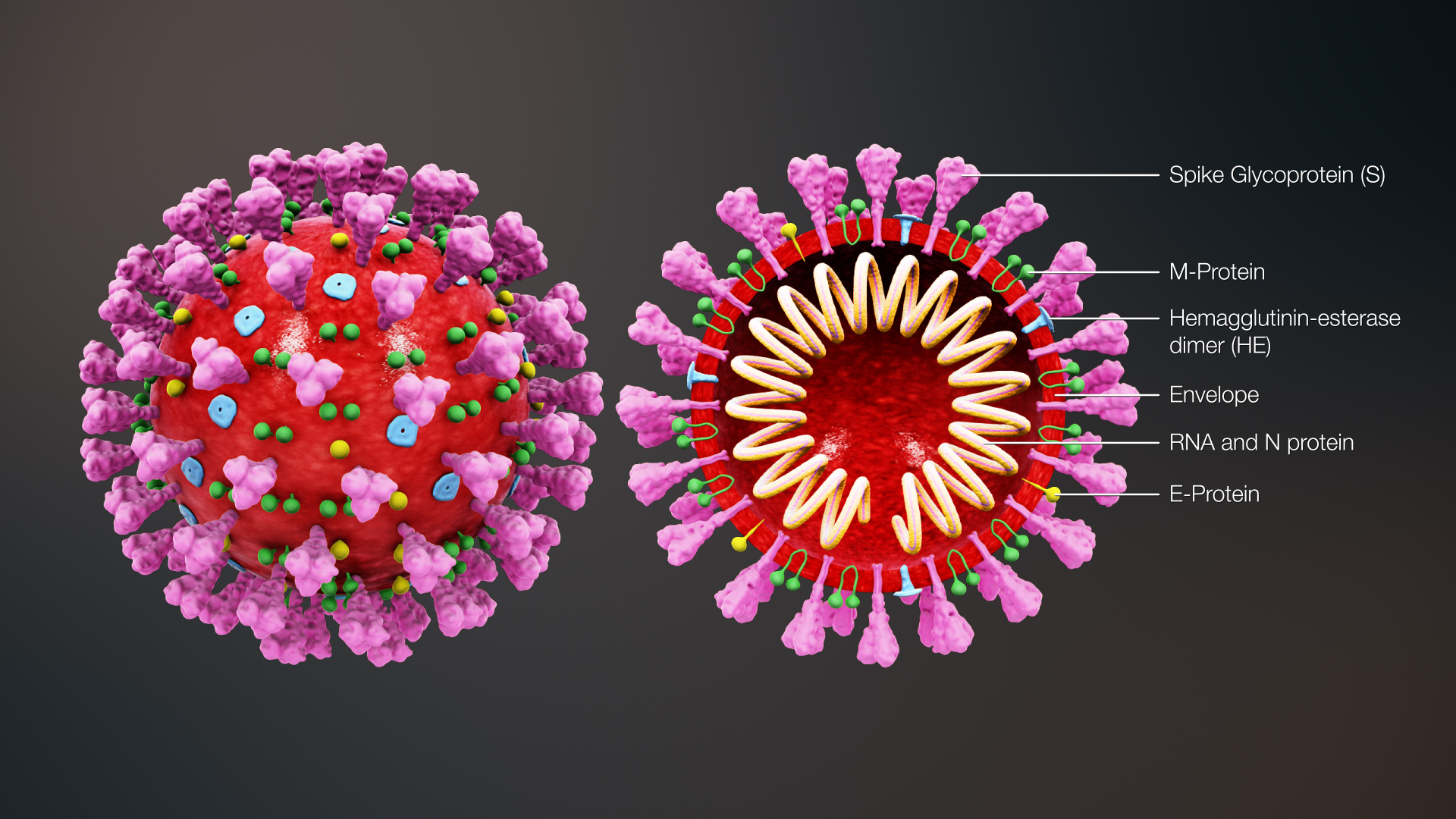
“As proteins also form part of the virus’s structure, we want to find a way to stop them using our cells to manufacture them – to prevent them from replicating,” says Professor Williams.

Health & Medicine
Q&A: How could COVID-19 drugs work and what’s out there?
“One hallmark of encapsulated viruses is that they harness the process of ‘glycosylation’ in the host cell. This is a key step in protein production where glycans, or sugar molecules, are added to newly assembled proteins.”
The sugar molecules help proteins fold into their correct 3D structure and they act as traffic signals to ensure the protein is delivered to where it is needed within the cell. Glycosylation is facilitated by various host cell enzymes that synthesize, trim, check and modify these sugar molecules.
Most proteins manufactured in the cell need glycans or sugars for folding ‘quality control’, and to provide directions on where in the cell to send the protein.
Human proteins usually undergo this quality control in a part of the cell called the endoplasmic reticulum. But a less commonly used pathway, and one preferred by certain proteins of encapsulated viruses, occurs within another part of the cell called the ‘Golgi apparatus.’
The Golgi is normally the cell’s postal service, shuttling proteins to the cell surface in a series of transport bubbles or ‘vesicles’. But for encapsulated viruses, some of the quality control that normally occurs in the endoplasmic reticulum occurs in the Golgi instead, and this preference could become the virus’ Achilles heel.
The human enzyme known as ‘MANEA’ (short for endo-α-1,2-mannosidase) is unusual as it performs sugar trimming steps in the Golgi. When appropriate trimming doesn’t occur, these proteins are marked as defective and discarded through waste disposal pathways.

Health & Medicine
Science, society and drug design
So the research team considered that MANEA could offer a new target for broad spectrum drug development against encapsulated viruses, by causing incorrect processing and triggering disposal of the essential viral proteins.
The key breakthrough in their study was producing enough of the MANNEA protein to study it, an effort that that took over five years’ work in Professor Davies’ lab.
The protein was then crystallised to study its 3D structure and understand which parts were crucial for the glycosylation process.
Once they had the structure, Professor Williams and his team were able to identify and develop a series of inhibitors to test which one best blocked MANEA’s activity.
The researchers were able to show binding between the inhibitors and MANEA, but the big question was do these inhibitors affect the replication of encapsulated viruses?
To test this, the team exposed human cell lines to bovine viral diarrhoea virus and then added the inhibitors to see what effect they had.

“We initially worked with bovine viral diarrhoea virus, because it is safe to work with, has established experimental tools and is a great model for encapsulated viruses, such as SARS-CoV-2,” explains Professor Davies.
Sciences & Technology
Go with the gut: Our symbiotic relationship with our intestinal bacteria
“Our team showed that inhibitors of MANEA that worked against the human mammalian enzyme, also worked to inhibit the replication and reduced the ability of BVDV to infect human cells.”
Having shown proof of concept in BVDV, they extended their work to another encapsulated virus, dengue virus, and demonstrated that the inhibitor reduced the replication of dengue virus too.
“It’s obviously timely. The work that we’ve done shows that this approach works for two encapsulated viruses, so it seems possible this could be a general mechanism that works against more encapsulated viruses, says Professor Williams.
“Because all these viruses use the same host pathway, a host-directed therapy has the potential to be a broad-spectrum anti-viral agent against all encapsulated viruses, potentially even on viruses we are yet to discover.”
Banner: Coronavirus infecting human cells. Getty Images