
Health & Medicine
Viruses on the wing

New research investigates whether Antarctic penguins have fewer viruses than other birds and if they act as a reservoir for viruses in birds globally - including the coronavirus
Published 15 April 2020
When I imagine Antarctica, I think of images of a completely isolated and hard-to-get-to place with lots of penguins, seals and, of course, Sir David Attenborough’s voice narrating in my brain.
But what we often forget is that the diversity of wildlife on the frozen continent also includes microorganisms.
For decades now, researchers have been interested in the viruses of Antarctic penguins in particular.

Studies of serology began in the 1970s.
This research compared antibodies to known viruses to try to find viruses that may be present in Antarctic penguins – serological experiments look at the immune response in blood, in this case the penguin’s response to viruses.
These early studies had great limitations, and were only able to understand whether penguins were infected with viruses we had previously catalogued, and these were mostly viruses that caused disease in poultry.

Health & Medicine
Viruses on the wing
As methods for studying viruses improved, researchers took to genetic methods for screening penguins, such as the now commonly used polymerase chain reaction or PCR.
Again, the large limitation here is that these methods rely on identifying only previously described viruses, not new species.
Things have changed dramatically in the last years with the advent of metagenomics; a method by which we can sequence the genomes of all the microorganisms in a sample. This enables us to randomly sequence a sample so that we are no longer limited to searching for only previously described viruses.
As a result, since 2012 or so, studies have found 15 different of viruses in Antarctic penguins.
These mostly comprise avian paramyxovirus virus (now called avian avulavirus) and influenza A viruses. As found in our previous research into penguins, these viruses do not seem to cause any disease in their hosts.
So our study set out to uncover the diversity of viruses in Antarctic penguins, and whether their remote homes means they are infected with fewer kinds of viruses compared to birds on highly connected continents.

To do this, we used a state-of-the-art method called metatranscriptomics that looks at all of the genes that are ‘switched on’ or expressed in the sample, providing an unbiased technique to find viruses.
Swabs were taken from the penguin cloacae– the opening of the digestive, reproductive and urinary tracts of birds. The RNA (a different type of nucleic acid to DNA) was then sequenced for each sample, this included all of the RNA of the birds, their microbiome and their viruses.
Many of the well-known viruses including flu and coronavirus have RNA as a genetic code.
With this method, we are able to describe new viruses, which is critical if we are to understand viruses in birds that don’t cause disease in them, but may have the ability to spill over into other species – including humans.

Health & Medicine
How our ‘avian athletes’ could spread influenza
We worked with collaborators at the University of Sydney and Universidad de Concepcion (Chile) to reveal virus communities of three species of Antarctic Penguins – Chinstrap Penguins, Gentoo Penguins, and Adelie Penguins on the Antarctic Peninsula.
This allowed us to assess whether these Antarctic penguins had fewer viruses than other birds and to understand the role of Antarctic penguins in acting as a reservoir for viruses in birds globally.
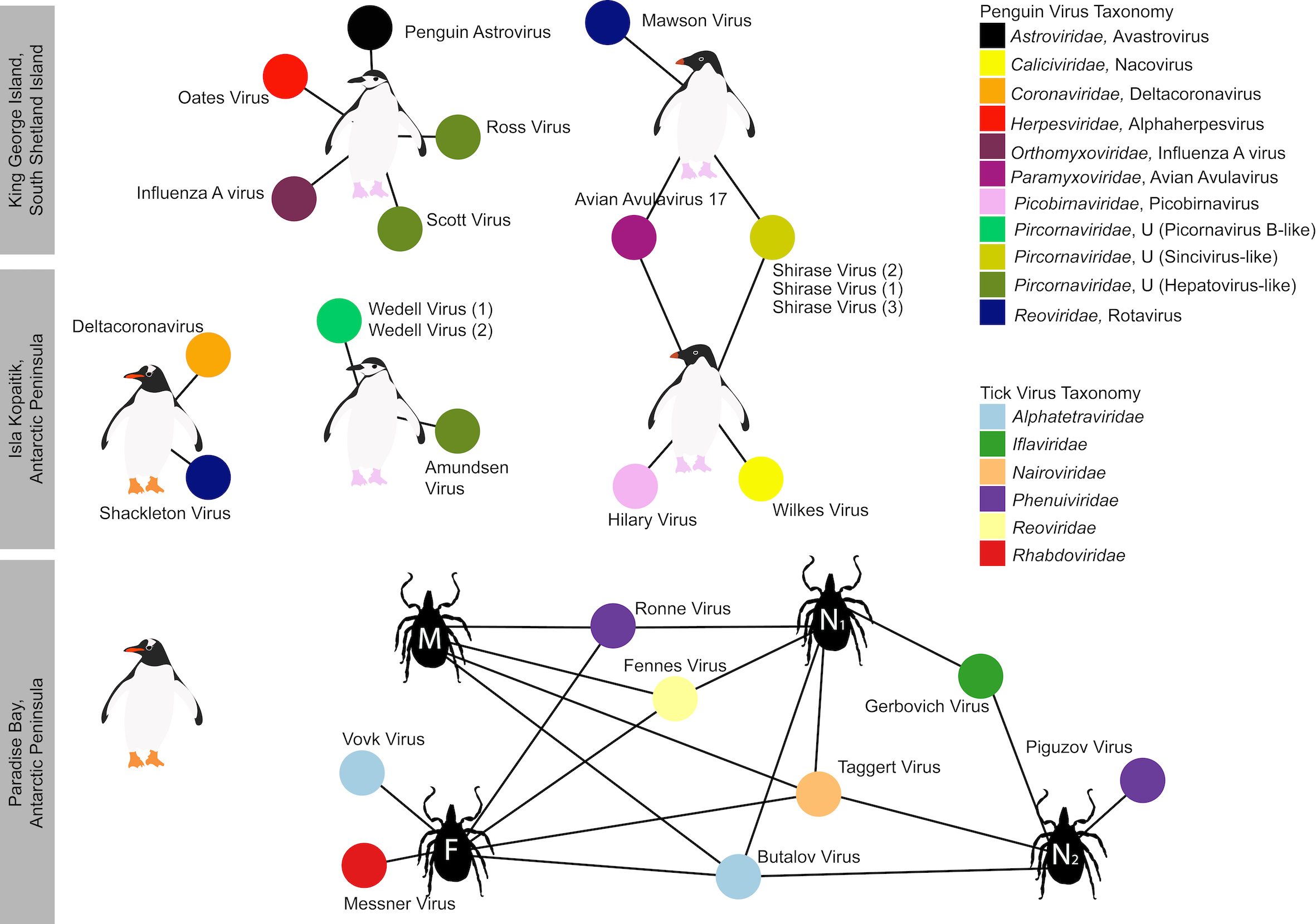
We found an amazing 107 viruses in total. From the six penguin sample pools (comprising a 60 samples from three species) we found 13 viruses that likely infect the penguins – essentially doubling the number of viruses known to infect Antarctic penguins.
Interestingly, when we use the same metatranscriptomic methods to find viruses in Australian wild birds, we tend to find a similar number of viruses per sample pool, really suggesting that these penguins feel a similar “pathogen pressure”.
This means that they are infected with a similar number and diversity of viruses as birds elsewhere in the world, despite the perceived isolation of Antarctica.
These viruses may arrive on the frozen continent through birds that have long distance migrations, such as Antarctic Tern; long distance wanders (as many seabirds are) or birds such as skuas and gulls, which move between the Antarctica peninsula and the southern reaches of Patagonia.
We found some viruses that have been described before in Antarctica, in addition to many new ones. For example, we also found a virus that matches closely to a very short virus sequence that was derived from penguins sampled in 2012 from the opposite side of Antarctica, along the coast of the Ross Sea.
This is important because it means that penguins are probably important hosts for these viruses, rather than just being occasionally infected, known as reservoir hosts.

Sciences & Technology
The invisible colours protecting birds from overheating
In one of the penguin species we also found a coronavirus.
Coronaviruses are actually rather common in birds, and have been found in wild birds wherever they have been tested. This implies that avian coronaviruses are globally distributed – from ducks in Sweden, to shorebirds in Australia and penguins in Antarctica.
This particular coronavirus that we found is in a group called the deltacoronaviruses, in comparison to SARS-CoV-2 (which causes COVID-19) which is a betacoronavirus. Deltacoronaviruses are mostly limited to birds, however, there is a virus found in pigs called Porcine coronavirus that is also in this group.
It may come as a surprise to many, but penguins do have ticks and these ticks do carry viruses that may infect penguins. We found eight viruses in ticks, of which, we believe two are viruses that may also infect the penguins.
One of these viruses, Taggart Virus, was first described on Macquarie Island in 1972 in ticks found in penguin colonies. This allows us to speculate that is may be a wildly distributed virus of penguins, but we would need to do more testing to figure that out.

Because of the powerful sequencing technology and the use of cloacal swabs, we also found 82 viruses in the penguin sample pools that are associated with their diet and microbiome.
These include invertebrate and fish viruses – these obviously don’t infect penguins, but they do infect the food they eat.
From just this small study, we found a large diversity of viruses, which is just the tip of the viral iceberg. With more sampling we anticipate to find more viruses.
Importantly, some of these new viruses will have the ability to spill-over into domestic animals or humans– which means it’s crucial to understand more about them.
Banner: Gentoo and Adelie penguins, Michelle Wille