
Sciences & Technology
The invasive fungus threatening Earth’s biodiversity

An international collaboration of researchers has found that frog diseases can be detected in environmental samples like soil and water
Published 16 September 2020
Detection of small amounts of DNA in environmental samples like water or soil is a new and exciting technology. Environmental DNA (eDNA) detection is a valuable conservation tool that can be used to identify and monitor imperilled or invasive species and even pathogens.
One such pathogen that is of global conservation concern is the fungus, Batrachochytrium dendrobatidis, Bd. Bd is a pathogen of conservation concern because it is a leading cause of frog declines around the world, by causing the disease chytridiomycosis.

Populations of many Australian frog species have declined, and some are even on the verge of extinction because of this disease, including the southern corroboree frog and the baw baw frog which are now classified as critically endangered.

Sciences & Technology
The invasive fungus threatening Earth’s biodiversity
Bd is thought to have spread out globally from Asia, causing massive declines around the world over the last few decades. Human global movement is a likely cause of this spread.
Frogs are intentionally moved internationally through the amphibian pet trade and even as food in many countries. And they can also be stealthy hitchhikers and easily travel internationally or interstate with the movement of produce and the plant trade.
By looking for a unique sequence of DNA from a pathogen, eDNA can be used to monitor disease outbreaks or identify disease introductions, which are essential steps in any conservation effort.
The traditional method of detecting disease in frogs is by catching them and swabbing them for the presence of the pathogen.
To test for Bd, scientists gently rub a medical grade swab, just like one used to test for strep throat, across the frog’s skin, focusing on the hands and feet where the infection tends to concentrate on the animal.
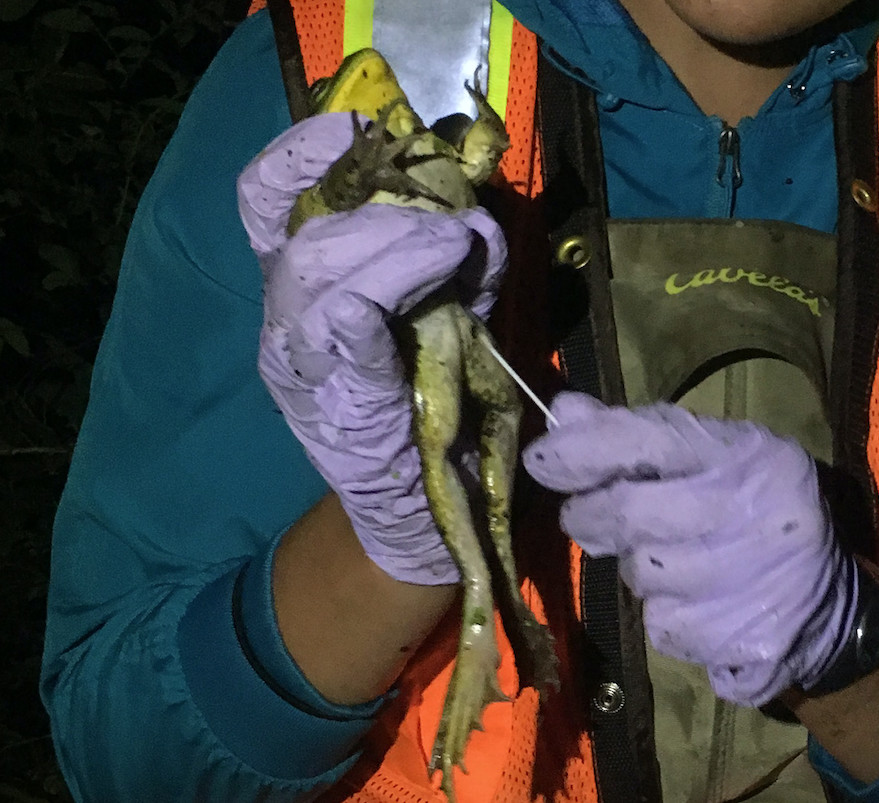

Catching animals can be challenging in difficult to access sites, or places with few or well-hidden frogs, such as the baw baw frogs that bury themselves deep in the mud of mountain gullies, or the southern corroboree frogs where there are only a handful of individuals left in the wild.

Environment
On the DNA trail of the platypus
In our recent study, we developed a method to detect Bd from both water and soil samples using lab-generated samples. Then we went into the field to see if we could potentially monitor disease using our eDNA method.
Our team of international scientists from the University of Melbourne and the University of Pittsburgh, USA, collected water and soil samples, and skin swabs from animals in multiple sites over six months of surveying.
The sites studied included highland streams, beaver ponds, swamps and seasonal forest pools in Pennsylvania and Louisiana USA. Some of these sites had over 10 species of amphibians.
Our results showed that eDNA techniques could detect the Bd fungal pathogen in the environment through both water filters and soil samples.
In fact, Bd detection in water samples was found to be just as good at detecting the pathogen in skin swab. While we were able to detect the pathogen in soil samples, it was not as accurate as water or skin swabs.
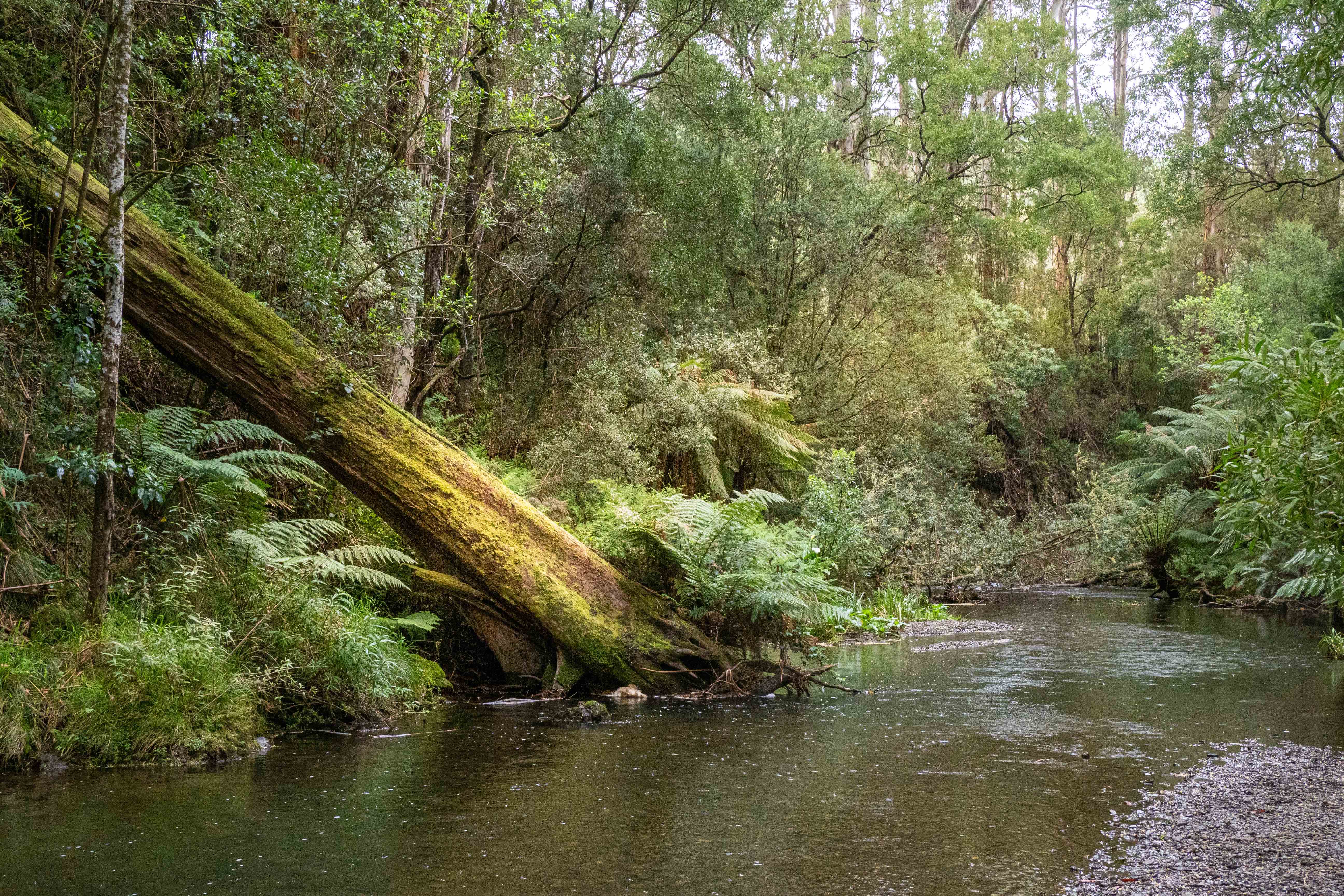

This is good news for studying pathogens that affect amphibians as there is plenty of water to test in their environments.

Sciences & Technology
Frogs and the City
The difference in the samples was that while we could detect the pathogen in water samples, the pathogen load estimates (the amount of pathogen detected) was more accurate in swab samples than from environmental samples.
This is somewhat surprising because chytrid fungi as a group are mostly soil dwellers, but makes sense because the infectious Bd zoospores are aquatic and move by swimming toward their next host. Also, with water filters, the larger volume of water filtered means a better rate of detection.
A key finding in this study was that there are limits to using eDNA for detection. We found that we can determine presence of the pathogen in the environment, particularly if there are many infected animals present.
However, eDNA is not a good method for determining the amount of pathogen in the environment; it is not as good as swabbing frogs for determining if the level of risk for high infection.

This is an important point, because while eDNA can be useful for monitoring, it should not replace the traditional methods of swabbing frogs. Instead eDNA can be used as a first step in monitoring sites more quickly and often less costly so that we can we can identify sites where more intensive monitoring is needed.
Sciences & Technology
The story in the bones of lizards and frogs
Another important limitation is that we can use eDNA to test for known pathogens, but it’s much harder to use to identify new or unknown pathogens.
By using eDNA, we can help monitor pathogen movement across the landscape, and it has the potential to be used to predict die offs, particularly in areas where the pathogen has not yet been introduced.
In this study we specifically investigated the devastating frog pathogen that causes chytridiomycosis.
However, this technique can be used more broadly for other pathogens affecting humans and agriculture. One example is that eDNA is currently being for bovine tuberculous to detect the risk of spillover from wildlife to agricultural cows.
Pathogen surveillance is important, especially when species are at risk of devastating declines.

The safest conservation measures are high biosecurity to ensure that pathogens are not moved around. But the second safest measures are to respond to pathogen invasions quickly. And therefore using a technique like eDNA can be important for those high risk locations, like Papua New Guinea.
Detecting pathogens and disease from environmental samples is exciting because it can be used for conservation of frogs around the world.
Our ultimate goal is to limit the introduction of pathogens into the environment and ensure that our unique amphibian populations can bounce back.
Banner: Southern corroboree frog/ Shutterstock