Sciences & Technology
Scientists decode sheep blowfly genome
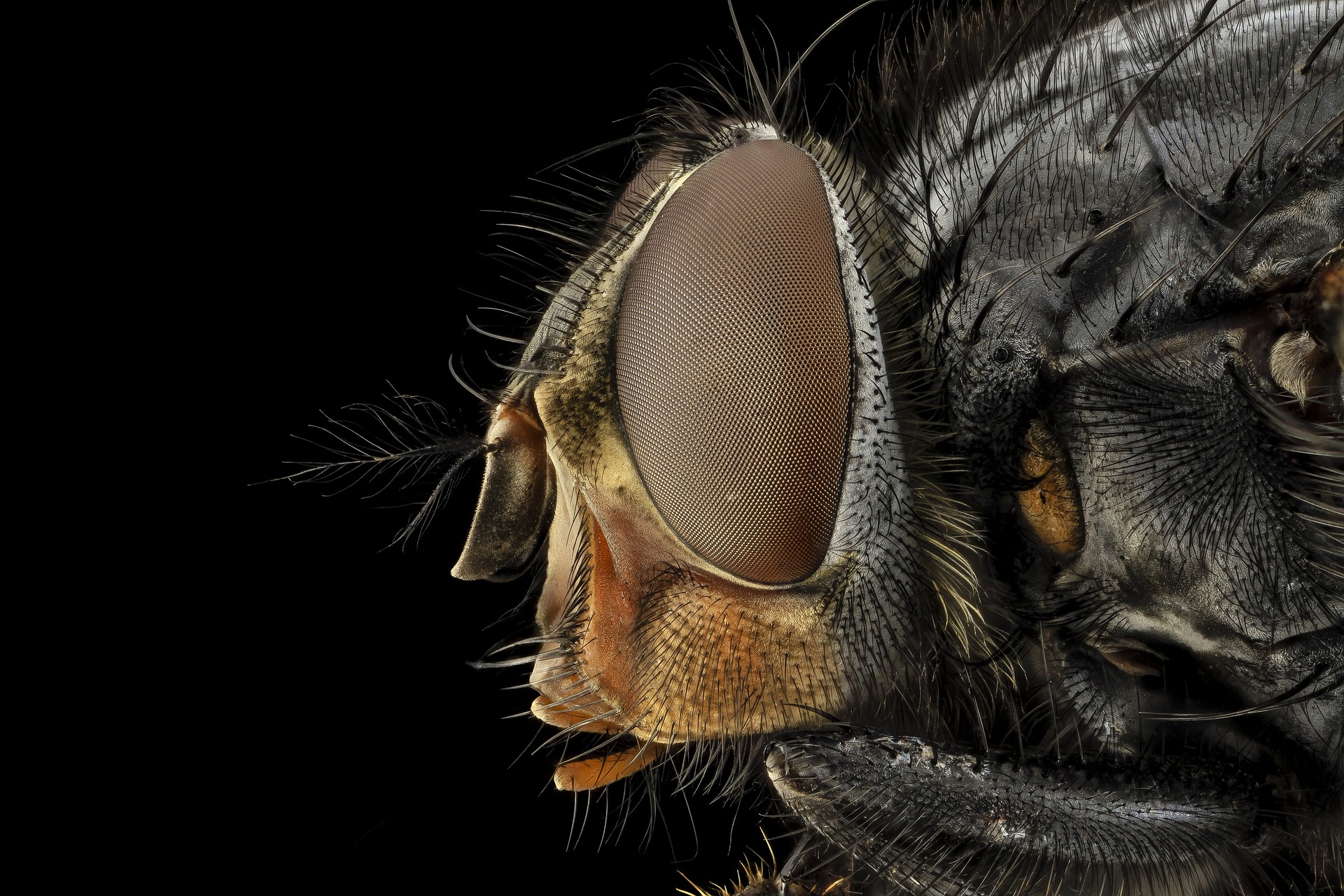

Melbourne researchers have been collecting blowfly samples from across Australia as part of the fight against flystrike
Published 7 June 2018
Opening the mail may not seem the most thrilling of tasks for a geneticist, but for a period of five months at the start of this year, our team was excited every time an Express Post envelope arrived at our Melbourne lab.
The envelopes contained sealed sample tubes of dead blowflies from sheep farms across Australia, carefully caught or picked off livestock and packaged by farmers keen to support our research.
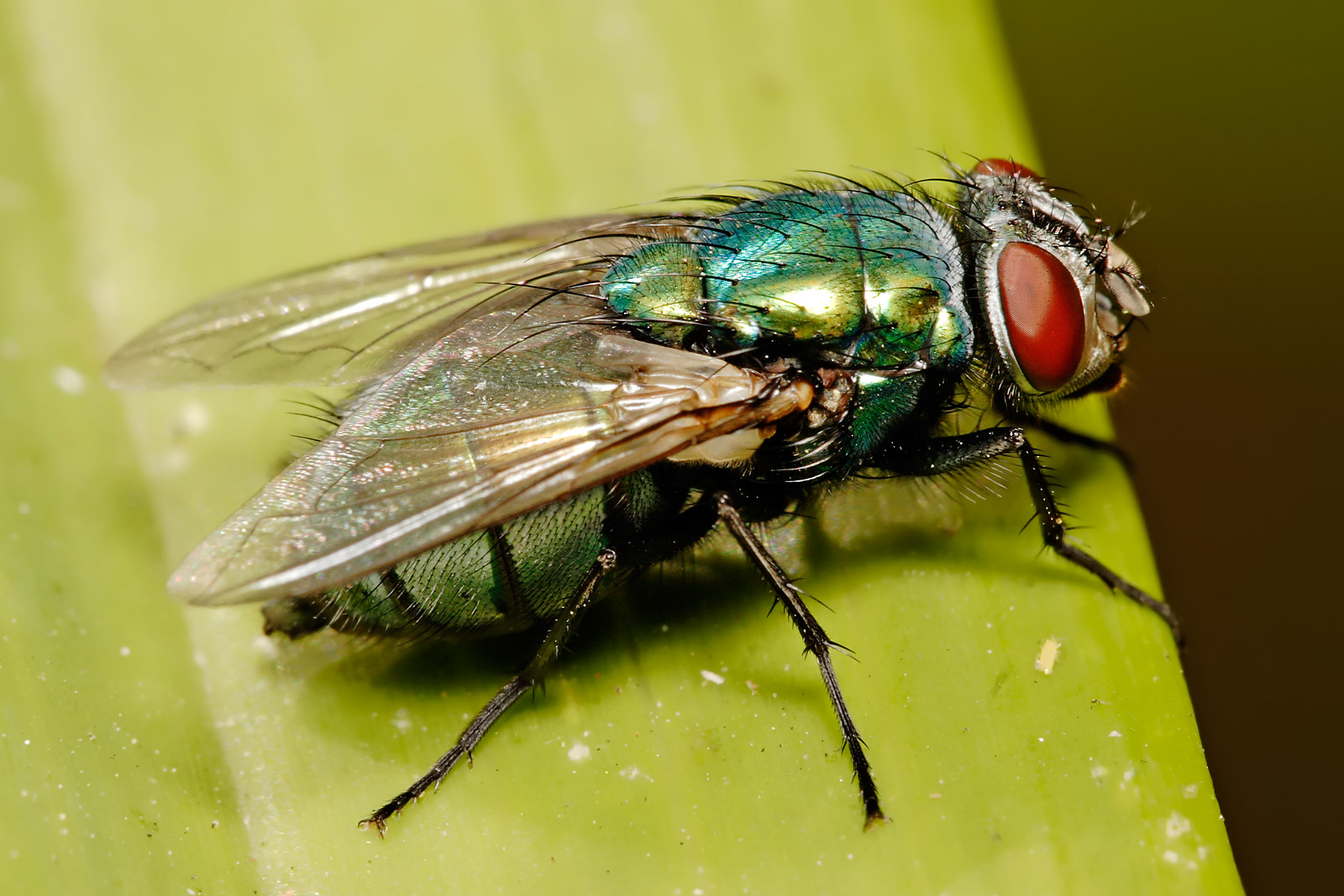

Not the most pleasant of packages, you might think, but vital for our work on a vaccine to beat flystrike.
Each year, blowfly infestation of sheep wool, skin and tissue, known as flystrike, results in $280 million losses to the wool industry, not to mention the suffering experienced by the animals themselves.
We are seeking to understand more about the blowfly’s genetics to design a vaccine against it. The hope is that by vaccinating the sheep, their buzzing enemies will be killed off before they cause harm.

Sciences & Technology
Scientists decode sheep blowfly genome
A particularly nasty disease, flystrike causes sheep considerable distress and can be fatal if left untreated. The flies lay their eggs in open wounds or matted wool, usually around the animal’s rear end where the flies are attracted to urine and faeces stuck in wool. When they hatch, the maggots migrate through the sheep’s wool to begin feeding on their flesh.
Having received 135 useable blowfly samples from over 17 locations in five states, we can now test what we’ve already learned about the early stages of the maggots’ development, to see if the genes we believe might be critical during this parasitic stage are conserved (have the same sequence) in blowfly populations across the country.
It’s taken a multidisciplinary team of scientists over six years to reach this stage, relying on some of the latest technologies in genetics.
We faced the somewhat daunting task of identifying the genes responsible for early maggot development, out of over 14,000 potential candidates in the Australian blowfly, (Lucilia cuprina) genome.

The first step was to map the fly’s full genome, a project led by Dr Anstead, published in 2015 after three years of work. Next, with the support of Australian Wool Innovation, we set about learning more about the early stages of how the maggots parasitize sheep, identifying the genes switched on during their first 24 hours of development.
We did this by analysing the ribonucleic acid (RNA) in the genes, which shed light on the genes and gene families expressed at the early development stage in the maggots, and to what extent. Then, using bioinformatic analyses (computer modelling to compare sequences of genes) to predict each gene’s likely function, we compared blowfly genes to databases of genes from other organisms.
Sciences & Technology
The tip of the CRISPR iceberg
Of most interest to us were the genes most likely to be exposed to their environment (and the sheep’s immune system).
Now we are looking to see if these genes are conserved in blowfly populations across the country. This is where our samples come in – we are sequencing their genomes to inspect whether the gene sequences we have identified are replicated in all our samples.
Once the DNA of our sample blowflies has been sequenced we will be able to validate our target genes and also determine the best type of sheep immune response to illicit.
It is important to understand how the blowfly would be affected by an immune response to our candidate genes. If they are vulnerable, we hope to harness that to develop a vaccine.

Sciences & Technology
Malaria’s dark secrets exposed by a simple glow
Sheep are good candidates for a vaccine to fight a parasite like the blowfly because their skin is very permeable. The maggots are quite prone to environmental stressors, so we are aiming to target a particular protein that will delay their development, for example, by inhibiting their feeding. This will be delivered through the sheep’s skin by targeting a specific immune response.
It’s a long and complicated road to a vaccine, but we will get there.
It’s in the sheep’s as well as the farmers’ best interests that new alternatives to prevent flystrike are developed, reducing the reliance on insecticides and other techniques that are currently the only effective options for controlling this pest.
This research has required a cross-disciplinary team to conduct the multiple faceted approach to finding and validating gene candidates, including members of the Batterham group from the School of BioSciences at the University of Melbourne and researchers from the Faculty of Veterinary and Agricultural Sciences (including Associate Professor Vern Bowles, Professor Robin Gasser and members of the Parasite Genetics and Genomics group).
Banner image: Pixabay