
Health & Medicine
Making the most of probiotics
Research into our gut microbiome – or our ‘second brain’ – has already produced vast amounts of data, but we are only scratching the surface of our own complex ecosystems
Published 14 January 2020
In recent years, our gut microbiome has expanded from the relative obscurity of a culture dish to become firmly established in popular culture.
Nicknamed the ‘second brain’ because it affects our mental health and behaviour – the gut microbiome is almost considered an organ of its own.
Most people now know that these microbes can affect our health, weight and mood, and that probiotics and prebiotics can improve our gut microbiome.
From the White House’s investment of US$121 million (around $A176 million) into the National Microbiome Initiative to commercial kits that we can use at home to interrogate our own biome – research into these microbial residents is growing.

Health & Medicine
Making the most of probiotics
However, our latest work at the University of Melbourne shows that we could make better use of the information we have on this complex ecosystem.
Designing experiments to understand the 1,500 trillion microorganisms residing on and in our body will always be complex, but we’ve found there are elements we can factor in to better understand our own ecosystem.
Beyond our gut biome, the full microbiome is a more extensive field of study looking at the entire collection of bacteria, fungi and viruses living inside and outside the human body.
These microbes create a highly dynamic and complex ecosystem that shapes our health.
Twenty-five years ago, our knowledge of the microbiome was limited to bacteria that could be grown or cultured in a dish. As most bacteria cannot be grown outside the body, our understanding of microbial interactions was limited.
With the advent of new techniques, we can now examine thousands of organisms in parallel by studying their DNA, without necessarily keeping them alive.

Initiatives like the Human Microbiome Project launched in 2007, followed by the Integrative Human Microbiome Project in 2014 have enabled researchers to study the microbial species that affect humans and their relationship to health and disease.
The emerging microbiome research field includes clinicians, microbiologists, bioinformaticians and statisticians. But despite this team effort, we are still only scratching the surface on how the microbiome is interacting with the host (that’s us) and shaping human health.
The amount of highly complex data means we still have a lot of work to do before we can identify microorganisms as accurate biomarkers for prognosis and diagnosis of disease.
Sciences & Technology
Go with the gut: Our symbiotic relationship with our intestinal bacteria
For robust microbiome studies, planning ahead is essential. The lifespan of a bacteria is about 12 hours, meaning the the microbiome is highly dynamic and sensitive to any changes in the environment.
So, if we want to study the effect of a specific diet on our microbiome, we need to choose exactly when to collect samples.
Researchers may not be aware of factors that can affect the quality of their data and the quality of the results from statistical analysis.
Statistician and geneticist Sir Ronald Fisher ruefully summarised this in 1938, when he said: “To consult the statistician after an experiment is finished is often merely to ask him to conduct a post-mortem examination. He can perhaps say what the experiment died of.”
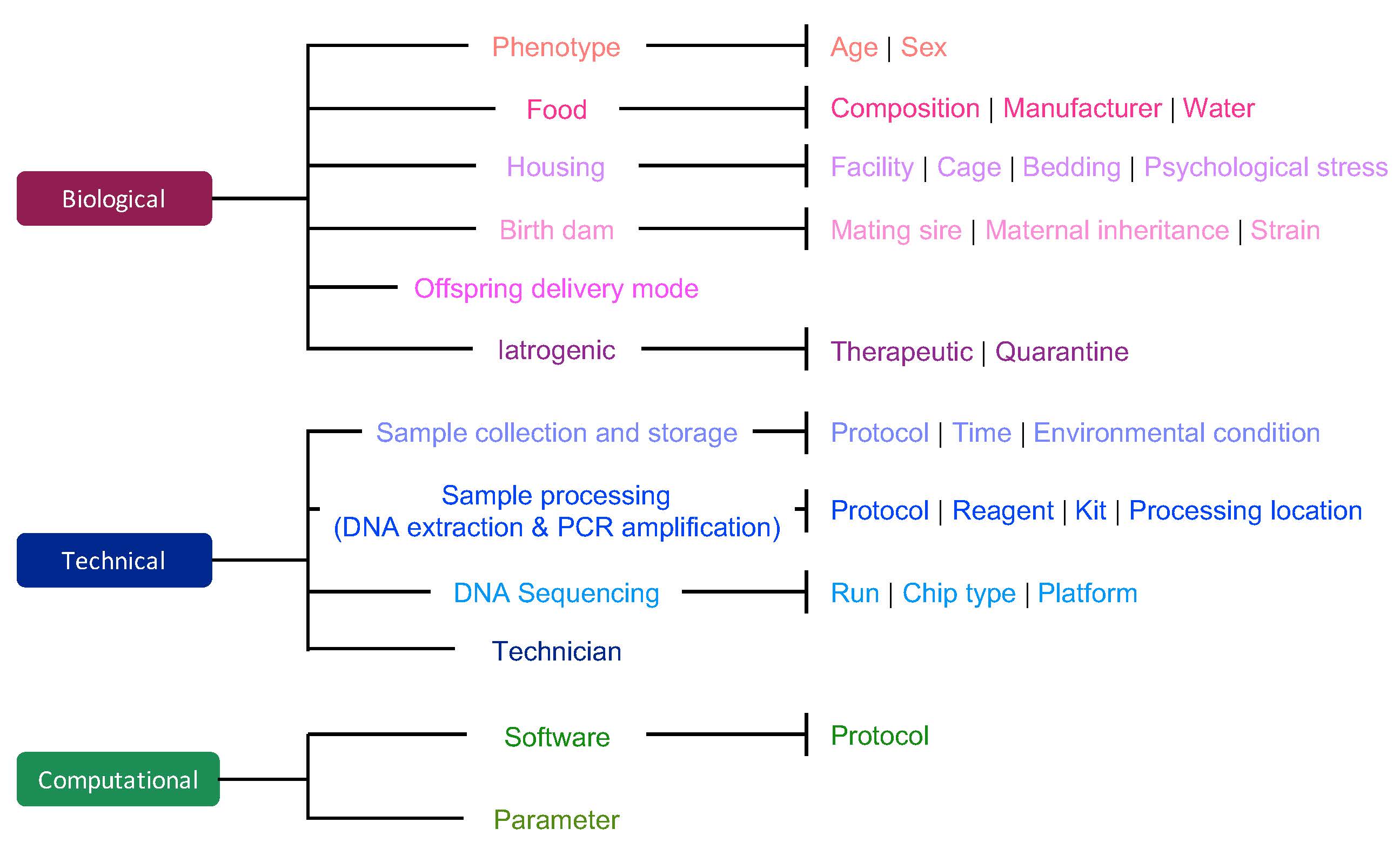
PhD candidate Yiwen Wang and I analysed several datasets to identify factors that may influence data quality.
These factors can be technical.
When studying microorganisms from faecal samples for example, the sequencing machine can only process a certain number of samples at once – known as ‘a sequencing run’.
From one run to another, we are likely to see differences in the microorganisms sequenced due to the slight differences in protocols or DNA extraction.

Health & Medicine
Antibiotics during pregnancy and the link to a baby’s immune system
Other factors can be computational.
The use of different software and algorithmic methods to process the data in a format ready for statistical analysis can generate different datasets.
Finally, biological factors also influence the microbiome.
Members of the same household are likely to share the same microbiome, as do mice hosted in the same cage for several weeks because diet and environment matter to our microbiome.
Other biological factors to affect the microbiome include the mode of birth delivery, like caesarian section or vaginal birth, as less beneficial and more potentially pathogenic bacteria initially colonise the newborn gut in caesarian section births.
Our methodological study with researchers from Université de Leval and National Research Institute of Science and Technology for Environment and Agriculture, both in France, confirm these findings are not anecdotal.
The contribution of statisticians in the microbiome field is essential to maximise the quality of results in these costly studies throughout a whole analysis cycle.
They can develop an analysis plan before the experiment to answer biological questions, while highlighting possible factors that may affect the data produced.

During and after analysis, statisticians educate and train those carrying out experiments in interpreting complex results, whether in silico – on the computer – in vitro or in vivo, which means in the lab or in the body.
Our study shows that specific microorganisms were more affected than others by technical or biological factors.

Health & Medicine
Chronic fatigue syndrome a kick in the guts
For example, specific bacteria can be found in increased abundance on Monday when we extract DNA from the first batch of samples, compared to Tuesday, when we process the second batch of samples.
These findings will certainly change the way we examine and analyse microbiome data.
Our next step is the development of a new statistical method that can directly deal with these factors.
This will reduce the ‘noise’ in analysis, so that data can be analysed efficiently to give better insight into the microbes that we all host.
Dr Lê Cao was recently awarded the Georgina Sweet Award for excellence in the area of Quantitative Biomedical Science.
Banner: Getty Images