
Environment
Helping fish fight their own battles

Advances in genomic and gene-editing technologies are giving aquaculture a much more precise way of making disease resistant fish and shellfish
Published 1 December 2020
Aquaculture – the farming of fish and shellfish, is the world’s fastest growing primary industry. It provides a healthy source of protein, oil and minerals for our rapidy expanding human population.
But viral and parasitic diseases remain a key challenge for aquaculture industries around the world.
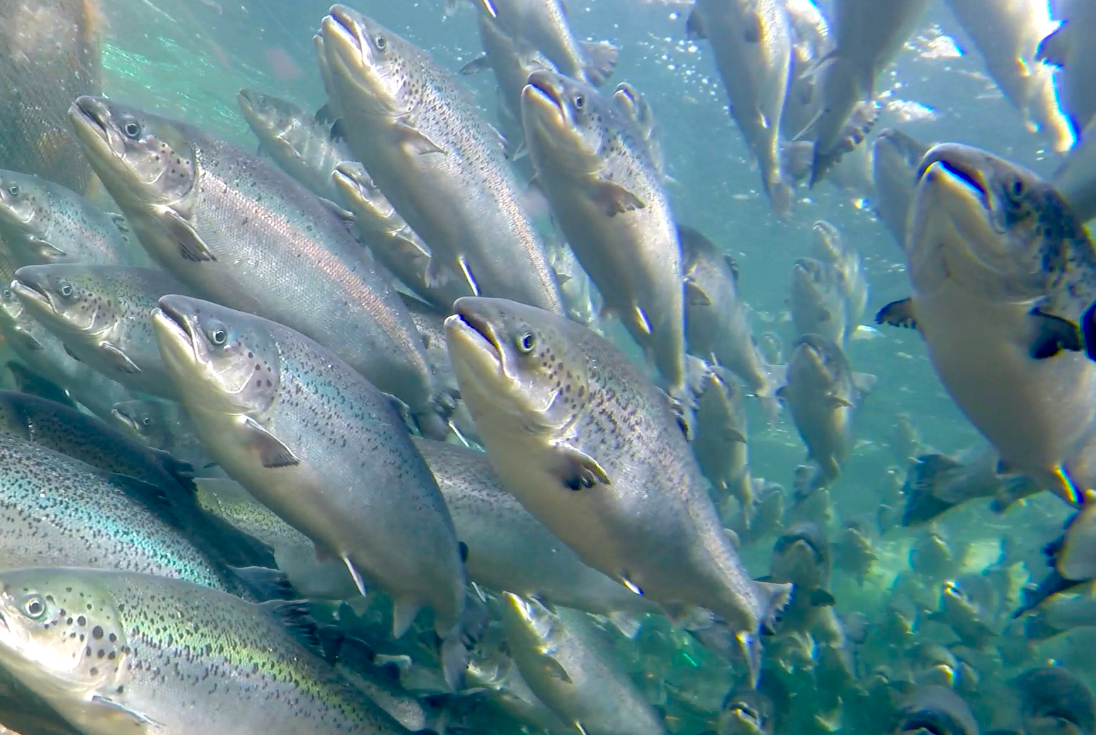

Our global experience this year with COVID-19 has highlighted that biosecurity is expensive and often limited in its protection against the spread of contagious disease.
Developing effective treatments – like vaccines – can be challenging, take many years and can be difficult to manufacture and deploy in order to effectively protect large populations.
The same is true for disease in aquaculture.
We know, however, that there is natural genetic variation in resistance for aquaculture’s most problematic viral and parasitic diseases.

Environment
Helping fish fight their own battles
So our research, some of which is published in the journal Scientific Reports, explores how this knowledge could be used in combination with the latest DNA technology advances, like genomic selection and CRISPR, to protect animals against disease.
White spot syndrome virus (WSSV) is a contagious and lethal disease in penaeid prawns that causes billions of dollars of losses globally. It can decimate whole prawn farms within a few days of infection and preventative measures have proven ineffective.
In late 2016, biosecurity efforts failed and WSSV disease was detected for the first time in Australia. The disease spread quickly from the initial farm to other farms and neighbouring wild populations in Queensland.
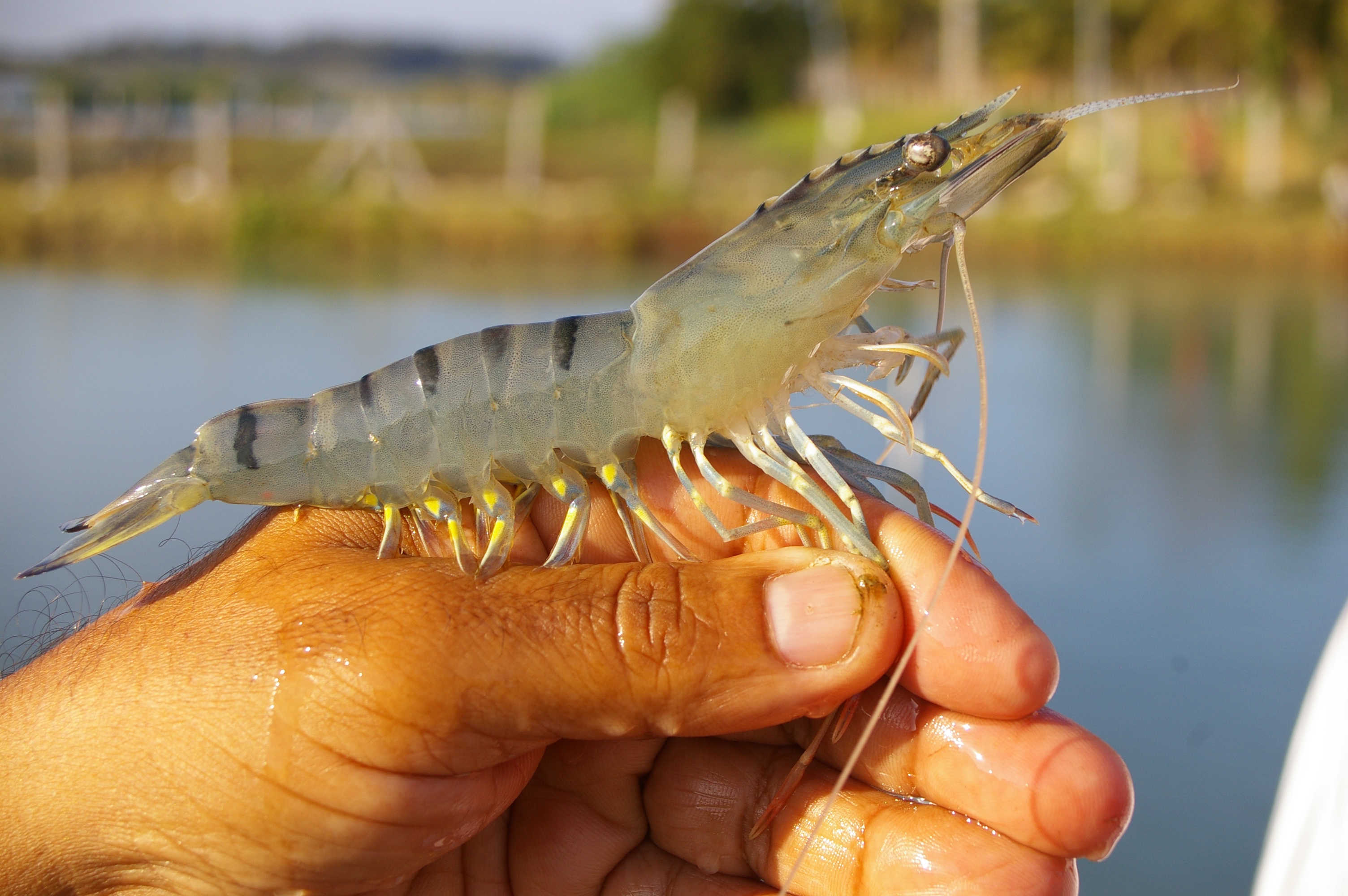

The outbreak prompted the largest aquatic animal disease emergency response ever undertaken in Queensland, costing $A4.4 million and depleting Australia’s supply of chlorine – around 3.8 million litres was used to clean ponds and water channels. It also resulted in temporary closure of the entire prawn farm industry, costing around $A400 million.
In April this year the disease resurfaced, re-infecting two farms. Like the rest of the world, Australia needs solutions that will help prawn farmers live with this virus.
Prawns and other crustaceans lack immunological memory, so vaccination for boosting the immune response has limited potential for protection against the virus.

Sciences & Technology
How data can keep fish and chips on the menu
Some animals are inherently better able to resist or tolerate the virus than others – however, we don’t really understand the specific mechanisms underlying these differences.
It is possible to breed animals with higher resistance to WSSV through conventional family selection, but progress has been slow. The industry desperately needs better, quicker solutions.
Our team of breeding and genetic scientists at the Norwegian Institute of Food, Fisheries and Aquaculture Research (Nofima) are working with a commercial partner, Benchmark Genetics, with funding from the Research Council of Norway, to boost the resistance of whiteleg shrimp (L. vannamei) against WSSV using genomic selection.
Instead of relying on pedigree relationships (family trees) to estimate the breeding value of individuals, our GenomResist project uses DNA sequence data to estimate the genomic relationships between individuals at tens-of-thousands of positions throughout the genome.
Some of the animals that are sequenced are challenged with WSSV, and response of these animals to infection, along with the genomic relationship data, can then be used to give us a much more precise way of predicting the disease resistance of potential breeding stock.
This is known as genomic selection.
A major advantage of genomic selection over traditional selective breeding for a trait like viral disease resistance is that it allows us to more accurately predict which breeders have the best overall resistance genotype without exposing the candidate breeders themselves to the disease.

Sciences & Technology
Farmed salmon are deaf – and now we know why
In an experiment using two shrimp populations developed by Benchmark Genetics Colombia, animals were randomly separated into two groups, one a test population which was challenged with the virus and the other a breeding population that was kept under high biosecurity.
By analysing variation across the genome in both populations, we could predict the genetic value of potential breeders.
We then selected and mated broodstock to produce two different populations of shrimp – one with high and the other with low genomic estimated breeding values. The survival of these two populations, and offspring from “randomly” mated parent stock, was compared in a virus challenge test.
We found that the average survival of shrimp families increased from 38 percent to 51 percent after only one generation of genomic selection for high white spot syndrome virus (WSSV) resistance.
Like the effect of vaccinating members of a population, high levels of immunity in the best populations can have a ‘herd immunity effect’ because these highly resistant animals will no longer infect other animals.
Benchmark Genetics now uses this tool to offer shrimp populations that can survive and produce in the presence of WSSV.

Environment
Meet Australia’s newest freshwater fish
Sea lice pose one of the biggest challenges facing the aquaculture industry worldwide. Sea lice wound and stress the fish. On top of this, the most effective methods of treatment (such as mechanical de-lousing) are also very stressful for the fish.
The cost to the Norwegian industry alone is more than US$550 million each year. Atlantic salmon are particularly susceptible to sea lice, while some species of Pacific salmon have very low susceptibility.
In a world-first project funded by the Norwegian Seafood Research Fund, our team from the University of Melbourne and Nofima, in collaboration with scientists from the UK, Canada and the USA, will work with large salmon breeding and production companies in Norway, using gene editing to find out which genes make salmon attractive as a host.
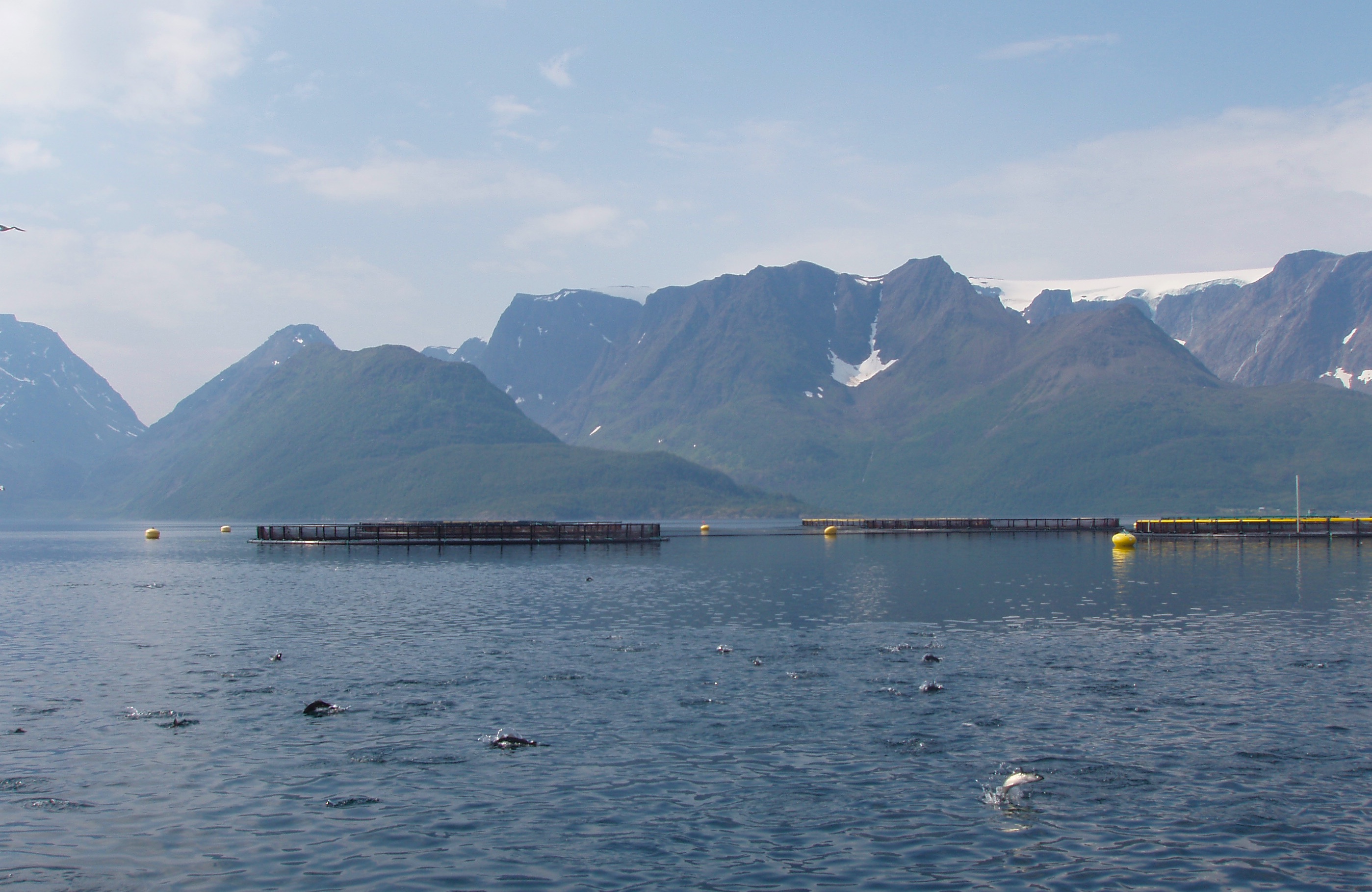

We are using the process for gene editing called CRISPR/Cas9 genetic scissors that won Emmanuelle Charpentier and Jennifer Doudna this year’s Nobel Prize in Chemistry.
If we can find the differences in the genetic code that cause lice to be attracted to Atlantic salmon, or that makes the skin of Pacific salmon unpleasant for sea lice to settle and develop, then we may be able to use that information to make Atlantic salmon resistant to sea lice.

Sciences & Technology
The ‘freak wave’ myth
Professor Tim Dempster and Associate Professor Ben Phillips at the University of Melbourne are also investigating the risk of sea lice adapting to the changes in the salmon, and how this would be best prevented.
The project will not make genetically edited fish available to the industry, but provide a thorough assessment of the potential for using this technology to improve the genetics of the fish, including possible effects on wild salmon populations.
Advances in our genetic knowledge and technologies are greatly increasing the accuracy and power of genetic improvement. But these new technologies also pose ethical issues for society that need to be openly debated and discussed.
Through these and other projects our team is working closely with industry and other researchers around the world to assess the potential for applying the latest sequencing and gene editing technologies to improve sustainability and welfare in aquaculture globally.
Banner: Getty Images